This website uses cookies so that we can provide you with the best user experience possible. Cookie information is stored in your browser and performs functions such as recognising you when you return to our website and helping our team to understand which sections of the website you find most interesting and useful.
The study relegates 'alternative splicing' as a source of protein production to a secondary level, which calls for rethinking the biology of higher organisms
The team argues that most human genes produce, against the prevailing opinion, a single dominant protein
This paper, published in the journal TIBS, also calls for review of theories on the origin and source of biological innovation
While the number of coding genes (those that produce proteins) in the human species has been consistently dwindling in recent years – the figures have fallen to fewer than 20,000-, it has been claimed that the dimension of the proteome, the element that executes the instructions in the genome, could be larger. This diversity of proteins has become one of the main sources of complexity in mammals, including the human species.
This theory could have an expiry date according to a study headed by the researcher Michael Tress from Alfonso Valencia’s group at the Spanish National Cancer Research Centre (CNIO), published today in the journal Trends in Biochemical Sciences (TIBS). According to the researchers, most genes produce, against the prevailing opinion, a single dominant protein. These results require the reassessment of the origin and source of biological innovation, which led to the emergence of primates 50 million years ago or to the development of the human brain, for example.
RESIZING THE HUMAN PROTEIN MAP
“The diminishing human genome” is how Valencia described the continuous corrections to the annotations of the human genome more than two years ago. Then, his team set the number of genes at around 19,000. Can something as complex as a human being be built from such a small number of genes?
Many researchers, sceptical about this issue, have turned their attention to the proteome as a possible source of biological innovation. Each gene can produce up to dozens or hundreds of RNAs, which result from combinations of various portions of a gene through alternative splicing. Then, the RNAs are translated into proteins. That is why alternative splicing has been identified as an important source of protein diversity.
In this chain of life, from the gene to RNA and from RNA to a protein, the authors of the paper realised there was a vast difference between the number of RNAs or transcripts, of the order of hundreds of thousands in humans, while the number of proteins, quantified experimentally, amounted to little more than 12,000. “The problem is the huge number of transcripts led us to assume there is a larger number of proteins, but the presence of all of them within the cells has never been demonstrated”, explains Michael Tress, principal investigator on the project.
“One gene, one protein, or one gene, several proteins?” the researchers ponder in the pages of the magazine. To answer this question, they conducted a comprehensive meta-analysis compiling data from eight large-scale experiments and from proteins or human peptide databases. The data analysed came from a wide range of tissues, cell lines and from different development stages.
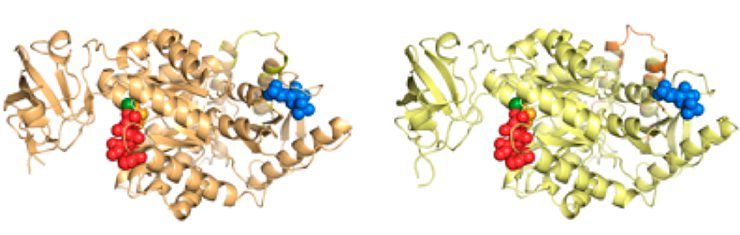
Pyruvate kinase variants, involved in glucose metabolism./ Michael Tress. CNIO
The results show that while there are many alternative variants of RNAs from a single gene, only a few genes (246, slightly more than 1 per cent of the human genome) presented clear evidence of producing more than one protein. “Most genes produce a single dominant protein. This tells us that alternative splicing is not essential for the complexity of the proteome”, explains Tress. According to the authors, when alternative splicing takes place it generates highly conserved proteins, with evolutionary origins that can go back more than 500 million years and with very subtle changes in their structure and function.
PREDICTING THE CONSEQUENCES OF DIFFERENT GENETIC VARIANTS
These observations may have significant implications in biomedicine, particularly in predicting the effects of genetic variants or mutations in the body. The team suggests that only the mutations in the DNA that have an impact on the dominant proteins will be detrimental.
Despite the limited evidence of alternative splicing in healthy cells, the situation is different with diseases such as cancer, in which this process plays a fundamental role in generating new forms of proteins with aberrant functions that compromise the viability of the organism.
Researchers are now pondering on the existence of all those RNAs for which no proteins have been detected and, therefore, for which we currently have no defined biological function. Could it be lost information? Useless information? Do they play new regulatory roles still to be discovered? If not from the alternative splicing, where does the proteome complexity come from? For now, there are questions that science is facing for which there are no answers.
This paper has been funded by the US National Institutes of Health (NIH).
Reference article
Alternative splicing may not be the key to proteome complexity. Michael L.Tress, Federico Abascal, Alfonso Valencia. TIBS (2016). DOI: 10.1016/j.tibs.2016.08.008
